Introduction
Label Distribution Learning is a novel machine learning paradigm. A label distribution covers a certain number of labels, representing the degree to which each label describes the instance. LDL is a general learning framework which includes both single-label and multi-label learning as its special cases.
Further details about LDL can be found in the following paper.
X. Geng. Label Distribution Learning. IEEE Transactions on Knowledge and Data Engineering (IEEE TKDE), 2016, 28(7): 1734-1748.
Our alogrithms can be used freely for academic, non-profit purposes. If you intend to use it for commercial development, please contact us.
In academic papers using our codes and data, the following references will be appreciated:
[1] X. Geng. Label Distribution Learning. IEEE Transactions on Knowledge and Data Engineering (IEEE TKDE), 2016, 28(7): 1734-1748.
[2] X. Geng, C. Yin, and Z.-H. Zhou. Facial Age Estimation by Learning from Label Distributions. IEEE Transactions on Pattern Analysis and Machine Intelligence (IEEE TPAMI), 2013, 35(10): 2401-2412.
Applications of LDL
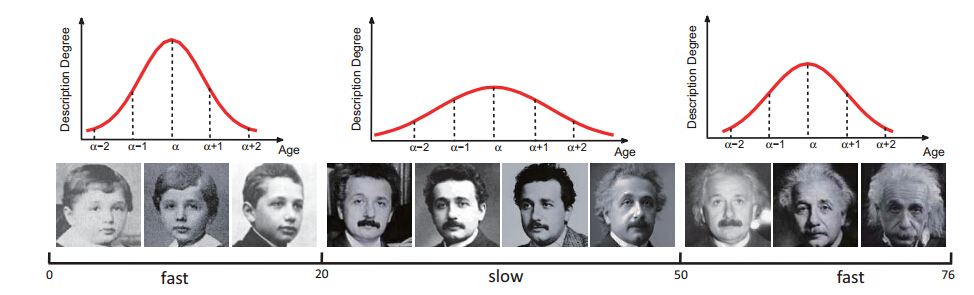 |
Facial Age Estimation
|
 |
Head Pose Estimation
|
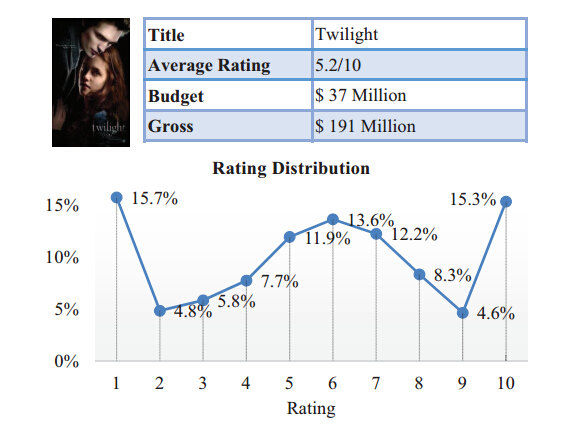 |
Pre-release Prediction of Movies
|
 |
Multi-label Ranking
|
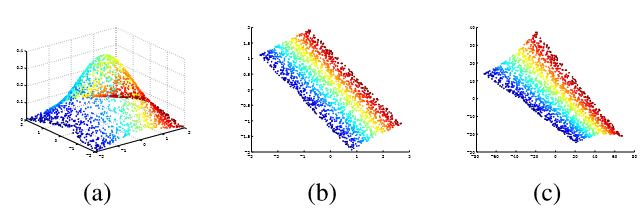 |
Multi-label Learning
|
 |
Crowd Counting
|
Matlab Code
We have implemented four LDL algorithms, namely IIS-LLD, BFGS-LLD, CPNN and LDSVR. To help you start working with LDL, we provide three demos (See iisllddemo.m, bfgsllddemo.m, cpnndemo.m, ldsverdemo.m) in this package.
Once downloaded, unzip the compressed folder to see the structure of LDL package. You'll see:
bfgsllddemo - The example of BFGSLLD algorithm.
bfgslldtrain - The training part of BFGSLLD algorithm.
bfgsprocess - Provide the target function and the gradient.
cpnn - The implementation of CPNN structure.
cpnndemo - The example of CPNN algorithm.
cpnnpredict - To predict the distribution by trained CPNN structure.
cpnntrain - The training part of CPNN algorithm.
drawdistribution - Draw the label distributions for comparision.
euclideandist - Calculate the average Euclidean distance between the predicted label
distribution and the real label distribution.
fidelity - Calculate the average fidelity between the predicted label
distribution and the real label distribution.
fminlbfgs - Finds a local minimum of a function of several variables.
iisllddemo - The example of IISLLD algorithm.
iislldtrain - The training part of IISLLD algorithm.
intersection - Calculate the average intersection between the predicted label
distribution and the real label distribution.
kernelmatrix - calculate the kernel matrix of vectors (samples) between two data matrices.
kldist - Calculate the average Kullback-Leibler divergence between the predicted label
distribution and the real label distribution.
ldsvrdemo - The example of LDSVR algorithm.
ldsvrpredict - To predict the distribution by trained LDSVR model.
ldsvrtrain - The training part of LDSVR algorithm.
ldsvrmsvr - The LDSVR's Multioutput SVR.
lldpredict - Calculate the predicted distribution of the instance X by weights.
sorensendist - Calculate the average Sorensen's distance between the predicted label
distribution and the real label distribution.
squaredxdist - Calculate the average squared x*x distance between the predicted label
distribution and the real label distribution.
Data Sets
We have collected 14 real-world LDL data sets. Here are some statistics about them:
| No. | Dataset | #Examples(n) | #Features(q) | #Labels(c) |
| 1 | Yeast-alpha | 2,465 | 24 | 18 |
| 2 | Yeast-cdc | 2,465 | 24 | 15 |
| 3 | Yeast-elu | 2,465 | 24 | 14 |
| 4 | Yeast-diau | 2,465 | 24 | 7 |
| 5 | Yeast-heat | 2,465 | 24 | 6 |
| 6 | Yeast-spo | 2,465 | 24 | 6 |
| 7 | Yeast-cold | 2,465 | 24 | 4 |
| 8 | Yeast-dtt | 2,465 | 24 | 4 |
| 9 | Yeast-spo5 | 2,465 | 24 | 3 |
| 10 | Yeast-spoem | 2,465 | 24 | 2 |
| 11 | Human Gene | 30,542 | 36 | 68 |
| 12 | Natural Scene | 2,000 | 294 | 9 |
| 13 | SBU_3DFE | 2,500 | 243 | 6 |
| 14 | Movie | 7,755 | 1,869 | 5 |
There are some other LDL datasets provided by other researchers, such as
| No. | Dataset |
| 15 | fbp5500 |
| 16 | RAF_ML |
| 17 | Twitter_ldl and Flickr_ldl |